be_polyelectrolyte¶
Polyelectrolyte with the RPA expression derived by Borue and Erukhimovich
Parameter |
Description |
Units |
Default value |
|---|---|---|---|
scale |
Scale factor or Volume fraction |
None |
1 |
background |
Source background |
cm-1 |
0.001 |
contrast_factor |
Contrast factor of the polymer |
barns |
10 |
bjerrum_length |
Bjerrum length |
Å |
7.1 |
virial_param |
Virial parameter |
Å3 |
12 |
monomer_length |
Monomer length |
Å |
10 |
salt_concentration |
Concentration of monovalent salt |
mol/L |
0 |
ionization_degree |
Degree of ionization |
None |
0.05 |
polymer_concentration |
Polymer molar concentration |
mol/L |
0.7 |
The returned value is scaled to units of cm-1 sr-1, absolute scale.
Note
Please read the Validation section below.
Definition This model calculates the structure factor of a polyelectrolyte solution with the RPA expression derived by Borue and Erukhimovich[1]. Note however that the fitting procedure here does not follow the notation in that reference as ‘s’ and ‘t’ are not decoupled. Instead the scattering intensity \(I(q)\) is calculated as
where
\(K\) is the contrast factor for the polymer which is defined differently than in other models and is given in barns where 1 \(barn = 10^{-24}\) \(cm^2\). \(K\) is defined as:
where:
\(b_p\) and \(b_s\) are sum of the scattering lengths of the atoms constituting the polymer monomer and the solvent molecules, respectively.
\(v_p\) and \(v_s\) are the partial molar volume of the polymer and the solvent, respectively.
\(L_b\) is the Bjerrum length (Å) - Note: This parameter needs to be kept constant for a given solvent and temperature!
\(h\) is the virial parameter (Å3) - Note: See [1] for the correct interpretation of this parameter. It incorporates second and third virial coefficients and can be negative.
\(b\) is the monomer length (Å).
\(C_s\) is the concentration of monovalent salt(1/Å3 - internally converted from mol/L).
\(\alpha\) is the degree of ionization (the ratio of charged monomers to the total number of monomers)
\(C_a\) is the polymer molar concentration (1/Å3 - internally converted from mol/L)
\(\text{background}\) is the incoherent background.
For 2D data the scattering intensity is calculated in the same way as 1D, where the \(\vec q\) vector is defined as
Validation
As of the last revision, this code is believed to be correct. However it needs further validation and should be used with caution at this time. The history of this code goes back to a 1998 implementation. It was recently noted that in that implementation, while both the polymer concentration and salt concentration were converted from experimental units of mol/L to more dimensionally useful units of 1/Å3, only the converted version of the polymer concentration was actually being used in the calculation while the unconverted salt concentration (still in apparent units of mol/L) was being used. This was carried through to Sasmodels as used for SasView 4.1 (though the line of code converting the salt concentration to the new units was removed somewhere along the line). Simple dimensional analysis of the calculation shows that the converted salt concentration should be used, which the original code suggests was the intention, so this has now been corrected (for SasView 4.2). Once better validation has been performed this note will be removed.
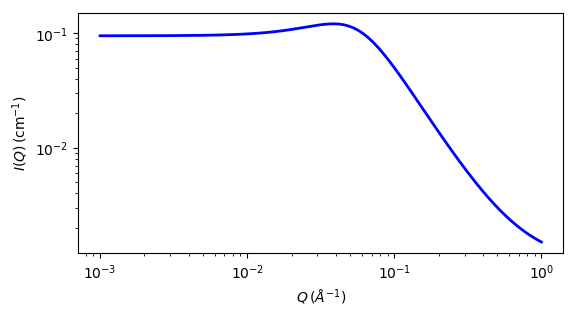
Fig. 98 1D plot corresponding to the default parameters of the model.¶
Source
References
For further details, see [2], [3], [4].
Authorship and Verification
Author: NIST IGOR/DANSE Date: pre 2010
Last Modified by: Paul Butler Date: September 25, 2018
Last Reviewed by: Paul Butler Date: September 25, 2018